David Chen
Postdoctoral researcher in Advanced Light Microscopy, Myers Group
Max Planck Institute of Cell Biology and Genetics, Dresden
Imaging biological systems across spatial scales is essential to understanding how tissues, organs, and organisms emerge from a mesoscopic collection of interacting molecules and cells. To this end, imaging tools must be able to both resolve subcellular structures with high spatial resolution and comprehensively map the tissue organization across large scales. Imaging mesoscopic samples imposes various challenges on microscopy. For example, thick and opaque samples require imaging methods capable of penetrating deep inside the tissue, large samples need high-speed imaging and efficient data handling, and quantitative analysis requires high-resolution images for accurate cellular and sub-cellular segmentation. To overcome these challenges, I have built a high-speed multi-beam two-photon microscope capable of imaging large fixed tissues–as big as one cubic centimeter–with sub-micron resolution routinely (a time-lapse video of the building process). I aim to use this novel technology to broaden our understanding of tissue architecture and self-organization across scales.
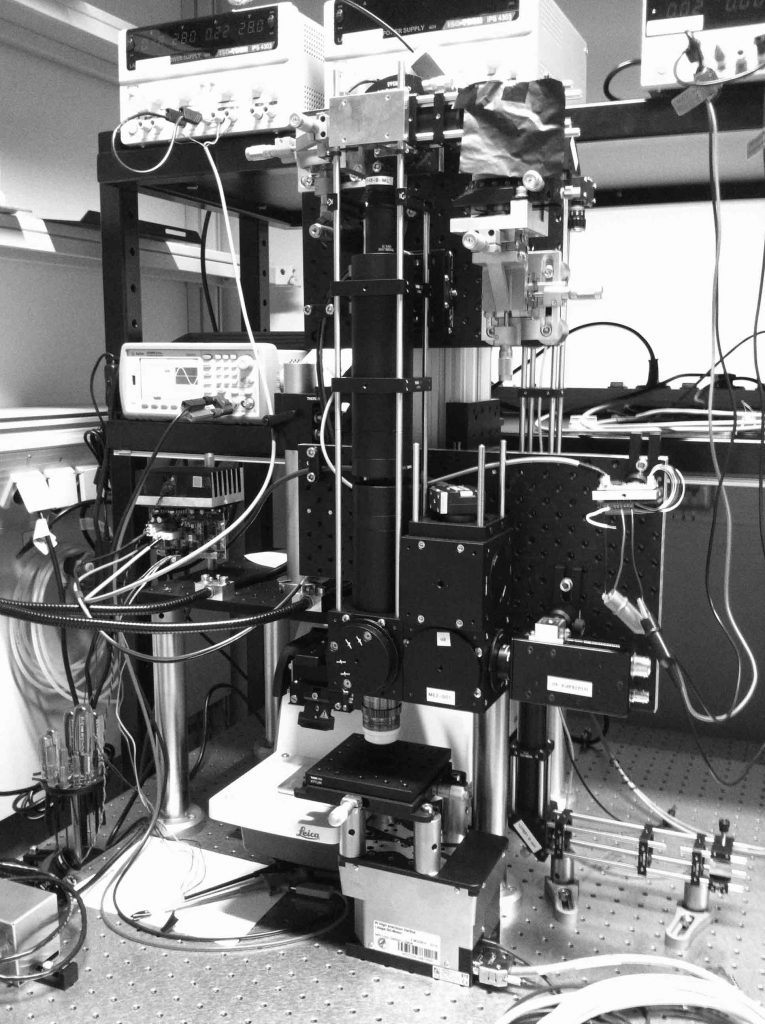
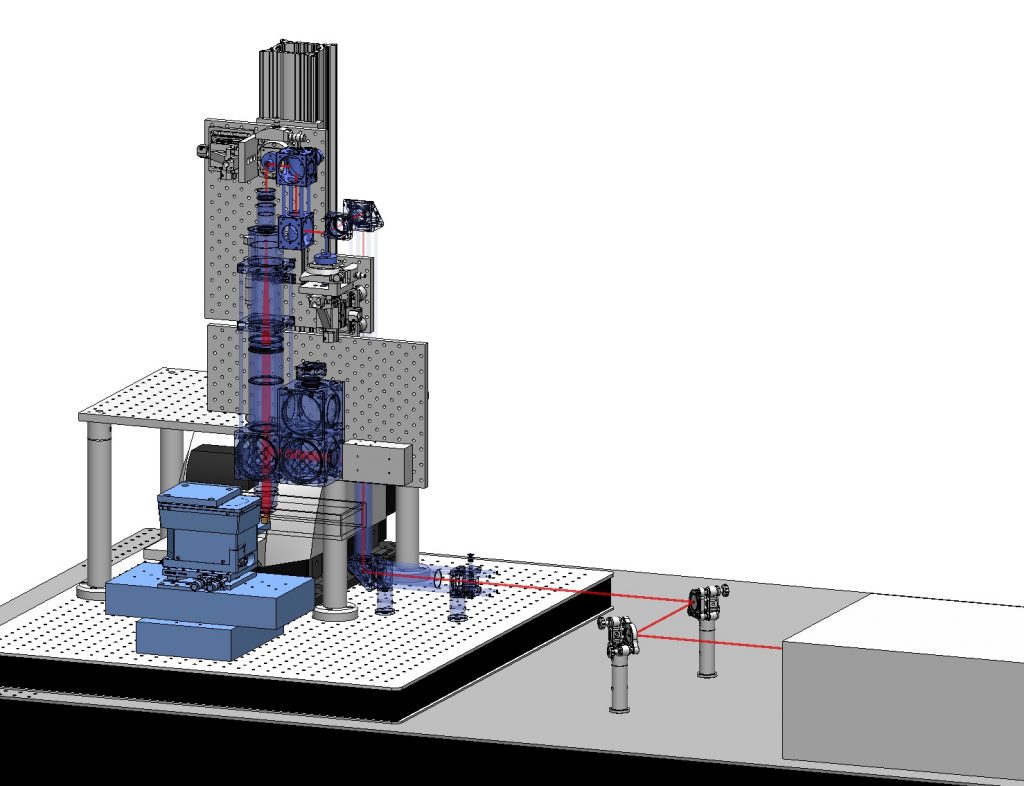

As part of this project, I have established a new laboratory for mesoscopic bioimaging at the MPI-CBG.
As a specific application of my microscope, I digitally reconstructed an entire mouse liver with multiple labels. I worked in close collaboration with biologists in the Zerial Lab at the MPI-CBG and theorists at the MPI-PKS to perform large-scale analytics on tissues (such as cell distribution, size, polarity, density, and connectivity) to aid the formulation of multi-scale models in systems biology.